A thorough
understanding of both the mechanism and the clinical application of drug
metabolism in the context of pharmacogenetics provides the clinician with the
greatest opportunity for identifying and determining the relevancy of a particular
interaction or adverse drug event. The table below is a summary of the main
genetic polymorphisms (or variations) of cytochrome P450 (CYP) 3A4 and if
known, the populations primarily affected, the specific genetic mutation, and
the impact of that mutation on enzyme activity.1-14 Compared to other CYP
enzymes involved in drug metabolism, it is well known that a large portion of
medications dependent on phase I metabolic pathways, will rely upon or be
substrates of CYP3A4.16 Medications substrates which are dependent on the
presence and/or functional activity of CYP3A4 may not be metabolized as
efficiently in the presence of one of these known genetic polymorphisms.
As such, these patients may experience exaggerated or unexpected pharmacologic
and/or side effects due to the higher concentrations of the medication
substrate present in the body.
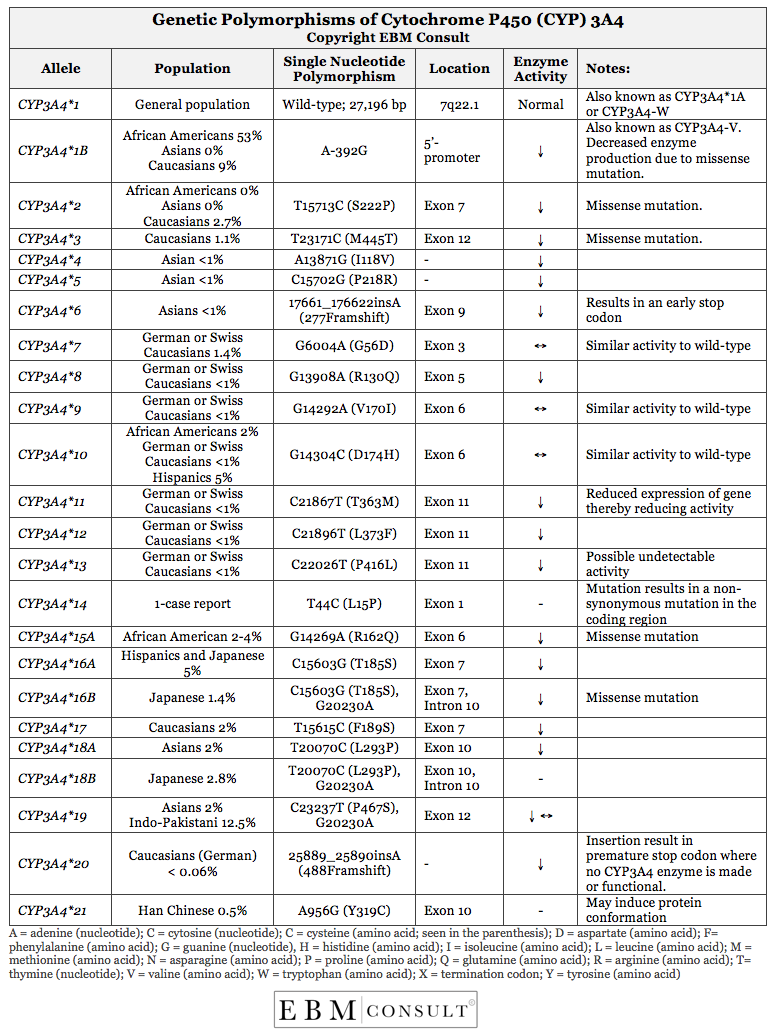
The important points to take away from this
publication include the following: 1) The majority of genetic polymorphisms to
the CYP3A4 gene result in decreased function and a few having no influence on
enzyme activity; 2) With the exception of the genetic polymorphism CYP3A4*1B,
CYP3A4*10 in Hispanics, and CYP3A4*19 in Indo-Pakistanis,
the proportion of patients without a genetic polymorphism appears low; and 3)
While some ethnicities are represented, there are still a large number of
patient populations where the impact of these genetic polymorphisms have not
been fully evaluated.
As a brief review, the genetic variations can
be interpreted by the location and/or type of mutation or defect in the genetic
code or sequence. For example, patients with the genetic
polymorphism CYP3A4*2 are known to have a single nucleotide
polymorphism at position 15,713 in the nucleotide sequence within exon 7 for
the gene that encodes for CYP3A4 enzyme. The nucleotide, thymine at
position 15,713 is changed to another nucleotide, cytosine. This single
change in the nucleotide changes the codon (3 nucleotide sequence) for the type
of amino acid placed at position 222 in the amino acid chain (once the gene
transcript has been translated by the ribosomes). Therefore, instead of a
serine (S; Ser) being placed at position 222 the amino acid, proline (P; Pro)
has replaced its position. This type of genetic variation is called a
missense mutation.
While there is still a need for clinical
research on the impact of genetic polymorphisms on drug efficacy and safety, we
do know that some of these variations have been associated with changes in drug
metabolism and/or elimination. For a list of medication substrates that
have the potential to be impacted by some of these genetic polymorphisms, we
recommend you go to the Drug Tables available
online.
References:
- Gonzalez FJ, Schmid BJ, Umeno M et al. Human P450PCN1: sequence,
chromosome localization, and direct evidence through cDNA expression
that P450PCN1 is nifedipine oxidase. DNA 1988;7:79-86.
- Rebbeck TR, Jaffe JM, Walker AH et al. Modification of clinical
presentation of prostate tumors by a novel genetic variant in CYP3A4. J
Natl Cancer Inst 1998;19:1225-9.
- Walker AH, Jaffe JM, Gunasegaram S et al. Characterization of an
allelic variant in the nifedipine-specific element of CYP3A4: ethnic
distribution and implications for prostate cancer risk. Mutations in
brief no. 191. Hum Mutat 1998;12:289.
- Sata F, Sapone A, Elizondo G et al. CYP3A4 allelic variants with
amino acid substitutions in exons 7 and 12: evidence for an allelic
variant with altered catalytic activity. Clin Pharmacol Ther
2000;67:48-56.
- Cavaco I, Gil JP, Gil-Berglund E et al. CYP3A4 and MDR1 alleles
in a Portuguese population. Clin Chem Lab Med 2003;41:1345-50.
- Sata F, Sapone A, Elizondo G et al. CYP3A4 allelic variants with
amino acid substitutions in exons 7 and 12: evidence for an allelic
variant with altered catalytic activity. Clin Pharmacol Ther
2000;67:48-56.
- Hsieh KP, Lin YY, Cheng CL et al. Novel mutations of CYP3A4 in Chinese. Drug Metab Dispos 2001;29:268-73.
- Van Schaik RH, de Wildt SN, Brosens R et al. The CYP3A4*3 allele: is it really rare? Clin Chem 2001;47:1104-6.
- Eiselt R, Domanski TL, Zibat A et al. Identification and
functional characterization of eight CYP3A4 protein variants.
Pharmacogenetics 2001;11:447-58.
- Lamba JK, Lin YS, Thummel K et al. Common allelic variants of
cytochrome P4503A4 and their prevalence in different populations.
Pharmacogenetics 2002;12:121-32.
- Dai D, Tang J, Rose R et al. Identification of variants of CYP3A4
and characterization of their abilities to metabolize testosterone and
chlorpyrifos. J Pharmcol Exp Ther 2001;299:825-31.
- Fukushima-Uesaka H, Saito Y, Watanabe H et al. Haplotypes of
CYP3A4 and their close linkage with CYP3A5 haplotypes in a Japanese
population. Human Mutat 2004;23:100.
- Lee SJ, Bell DA, Coulter SJ et al. Recombinant CYP3A4*17 is
defective in metabolizing the hypertensive drug nifedipine, and the
CYP3A4*17 allele may occur on the same chromosome as CYP3A5*3,
representing a new putative defective CYP3A haplotype. J Pharmacol Exp
Ther 2005;313:302-9.
- Westlind-Johnsson A, Hermann R, Huennemeyer A et al.
Identification and characterization of CYP3A4*20, a novel rare CYP3A4
allele without functional activity. Clin Pharmacol Ther
2006;79:339-49.
- Zhou Q, Yu X, Shu C et al. Analysis of CYP3A4 genetic polymorphisms in Han Chinese. J Hum Genet 2011;[Epub ahead of print].
- Rendic S, Ci Carlo FJ. Human cytochrome P450 enzymes: a status
report summarizing their reactions, substrates, inducers, and
inhibitors. Drug Metab Rev 1997;29:413-580.